March 2026 Research Highlights: Portal Powers
CAR-T Cell Engineering Directly from Whole Blood
Mechanoporation selectively permeabilized immune cells within whole blood while leaving other blood components (RBCs, platelets) unaffected, enabling simultaneous delivery of two RNA payloads to T cells without any upstream processing.
.png)
Experimental Setup
- Cell source: Unfractionated human whole blood
- Cargo: Circular RNAs encoding CD19 CAR and membrane-bound IL-2 (mbIL-2), co-delivered in a single step
- Readout: Flow cytometry at 40h post-boost for CAR and mbIL-2 surface expression on T cells within whole blood
Key Results:
- Boosted condition: 52.9% double-positive (IL-2+ and CD19 CAR+), 15.7% IL-2+/CAR-, 2.29% CAR+/IL-2-, 29.0% double-negative
- Control condition: 96.1% double-negative, 0.17% IL-2+/CAR-, 0.045% double-positive, 3.66% CAR+/IL-2-
- 52.9% double-positive in boosted vs. 0.045% in control
-1.png?width=1200&length=1200&name=image%202%20(2)-1.png)
Enabling Lentivirus Infection on CD19+ Population
CD19+ B cells isolated from PBMCs are resistant to standard lentiviral transduction. In this experiment, mechanoporation was used to pre-deliver a circular RNA coding for a transduction enhancer (TE) before viral exposure.
.png)
Experimental Setup
- Cell type: Primary human CD19+ B cells isolated from PBMCs
- Cargo: Circular RNA coding for a transduction enhancer (TE)
- Follow-up: 3-day incubation with GFP lentivirus post-mechanoporation
- Readout: Flow cytometry for TE expression and GFP+ transduction efficiency
Key Results:
- TE Expression: No Virus Control ~0%, No Virus Boost approximately 75%, Virus Control ~0%, Virus Boost ~60%
- GFP Expression: No Virus Control ~13%, No Virus Boost ~2%, Virus Control ~13%, Virus Boost ~47%
- Lentiviral transduction reached ~47% GFP+ in boosted cells vs. ~13% in unboosted controls (3.6-fold increase)
These data demonstrate that mechanoporation can function as a complement to existing viral workflows. By pre-delivering enhancer molecules that increase cellular permissiveness to viral entry, this approach extends lentiviral transduction to cell types that have historically been difficult to transduce.

CD19+ cells were isolated from PBMCs and mechanoporated with a circular RNA coding for a transduction enhancer (TE). The boosted cells were then incubated with a GFP lentivirus for 3 days. Surface receptor and GFP expression were assessed using flow cytometry after incubation period
Intracellular Protein Detection in Live Cells by Delivering Antibodies

Simultaneous intracellular delivery of 4 antibodies:
- Primary antibodies targeting 1) BTK and 2) p-BTK
- Lumit Secondary antibodies containing 3) LgBiT and 4) SmBiT
-
Luminescence occurs when LgBiT and SmBiT antibodies bind in proximity
Experimental Setup
- Cell types: Ramos B cell line and primary B cells
- Stimulation: Pervanadate treatment to induce BTK phosphorylation
- Cargo: 4 antibodies delivered simultaneously - primary antibodies targeting BTK and p-BTK, plus Lumit secondary antibodies containing LgBiT and SmBiT
- Mechanism: Luminescence generated when LgBiT and SmBiT antibodies bind in proximity to phosphorylated BTK
- Readout: Luminescence on GloMAX plate reader
Key Results:
- Ramos B cell line: Portal live-cell delivery produced ~1,100,000 luminescence units (pervanadate-stimulated) vs. ~1,350,000 for lysate, with minimal signal in no-cargo controls
- Change in p-BTK level is comparable between the lysate and intracellular live-cell assay
- Primary B cells: Experiment yielded comparable results between lysate and Portal-enabled intracellular live-cell assay
- All assays performed in intact, living cells - no lysis required
By bypassing lysis, this approach preserves native cellular context. Change in p-BTK level is comparable between the lysate and intracellular live-cell assay.
-1.png?width=1200&length=1200&name=Group%204%20(3)-1.png)
-1.png?width=1200&length=1200&name=Group%203%20(4)-1.png)
pBTK luminescence in primary activated B cells comparing Lysate and Portal. Lysate: ~200,000 untreated, ~700,000 stimulated. Portal: ~60,000 untreated, ~140,000 stimulated. Comparable fold change in p-BTK between methods, confirming live-cell antibody detection works in primary cells.
High-Throughput CRISPR Editing of Unstimulated T Cells
B2M RNP was pre-loaded into 96-well plate wells. T cells were mechanically porated through an in-line Portal cartridge during automated dispensing, with immediate RNP uptake upon entering wells.
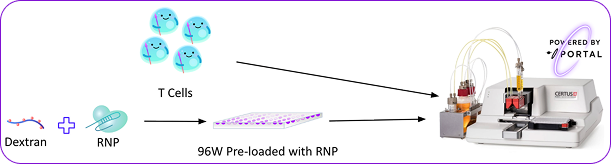
Experimental Setup
- Cell type: Unstimulated T cells isolated from a buffy coat (no pre-activation)
- Cargo: CRISPR RNPs targeting B2M + fluorescent dextran tracer, pre-loaded in 96-well plates
- Integration: Galaxy-i cartridge integrated into the Nnano Certus Flex (96-well plate format)
- Readout: Flow cytometry 3 days after delivery for viability, dextran uptake, and B2M surface expression
Key Results:
- 74% B2M knockout (B2M-negative by antibody staining), averaged across all wells after initial ramp-up stabilization (rows A-B)
- 80% dextran-positive cells, indicating high delivery consistency
- 72% viability at Day 3 (vs. ~83% untreated baseline)
Transcriptional impact: Mechanoporation produces fewer than 10 gene expression changes (>2-fold, p<0.05) compared to more than 100 for electroporation, preserving baseline T cell transcriptional profiles during the editing process.
Multi-target editing: Separate research scale experiments have demonstrated triple gene knockout (B2M + PD-1 + TRAC) with 40-80% editing efficiency across guides in primary T cells, using sequential mechanoporation.

B2M knockout (purple, avg 74%) and dextran delivery (cyan, avg 80%) across dispense number. Both metrics stabilize after ramp-up zone (rows A-B) with <5% well-to-well CV, demonstrating consistent high-throughput gene editing.

% live T cells across ~100 dispenses through Certus Flex system. Average viability 72% vs. ~83% untreated baseline. Gray "ramp-up zone" (rows A-B) shows initial stabilization before consistent performance.
High Throughput mRNA Screening in Primary Immune Cells
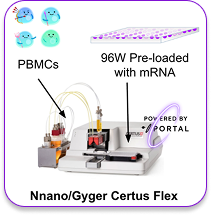
PBMCs were boosted using a Galaxy-i cartridge integrated into the Nnano Certus Flex, dispensing into 96-well plates pre-loaded with 27 different GFP mRNA constructs plus a positive control. GFP expression was measured at 20h post-boost by flow cytometry, with GFP MFI normalized to dextran delivery per well.
Experimental Setup
- Cell source: Human PBMCs
- Platform: Galaxy-i cartridge + Nnano Certus Flex, 96-well plate format
- Cargo: 27 different GFP mRNA constructs + positive control, pre-loaded in wells
- Readout: Flow cytometry at 20h post-boost; GFP MFI normalized to dextran delivery per well
Key Results
- 27 mRNA constructs screened in a single run using mechanoporation into PBMCs
- Wide range of GFP expression across constructs: Construct 22 achieved the highest GFP MFI (~700), while other constructs ranged broadly down to near-zero
- GFP MFI normalized to dextran delivery to account for well-to-well delivery variation
- Flow cytometry histograms confirm distinct expression profiles across constructs (No mRNA, Construct 8, 16, 5, 22)
This approach enables rapid ranking of mRNA construct performance in primary immune cells, screening dozens of candidates in a single 96-well plate run.

GFP MFI across 20 mRNA constructs ranked by expression level, with construct 22 highest (~700) and no-mRNA control near zero. Inset: flow cytometry histograms for selected constructs showing expression distribution.